METABOLOMICS AND ITS POTENTIAL TO UNDERSTAND THE MODE OF ACTION OF PLANT BIOSTIMULANTS
By: Luigi Lucini, Catholic University of Piacenza, Italy – email: luigi.lucini@unicatt.it

Although plenty of literature is supporting the benefits related to the use of biostimulants, the molecular mechanisms underlying such positive effects are still poorly elucidated. Nonetheless, the comprehension of the mode of action through which biostimulants exert their activity can provide useful insights to better define the target(s) in terms of crops, claims, agricultural practices and timing of application. Furthermore, recent regulation adopted by EU suggest the opportunity of understanding the crop processes affected by the biostimulant in order to support the desired claims at the regulatory level.
With this premise, we can point out that addressing the above-mentioned request is a critical challenge. This is due, mainly, to two reasons: (i) the complexity of plant biochemical processes, some of them not yet fully elucidated; (ii) the variability in genotypic response, rather than from cultivar to cultivar, and the pivotal modelling role exerted by both environment and agronomic practices.
Being anchored to soil, plants had to develop a huge diversity of biochemical processes to respond to both abiotic and biotic stresses; plant biostimulants often target/modulate these processes. Several approaches can be used to shed light onto the biochemical variations imposed by external factors such as the use of biostimulants on crops. Among them, untargeted approaches are to be recommended because they do not require a priori hypotheses, thus being amenable for complex scenarios. Omic sciences (e.g. genomics, transcriptomics, proteomics and metabolomics) refer to the profile of genes, transcripts, proteins and metabolites in a biological system and fall within untargeted approaches. Among the different omics, metabolomic is the closest to genotype and has been efficiently proposed to investigate the mode of action of plant biostimulants. Indeed, together with modulation of gene expression we can have post-translational modification of proteins, finally resulting in the actual metabolite profile. Recent advances in analytical equipment like high-resolution mass spectrometry, databases availability and data management (including multivariate statistical analysis and bioinformatic tools for data interpretation) have boosted the adoption of metabolomics in agricultural sciences. Consequently, differences in metabolomic signatures have been proposed to identify the plant processes affected by external factors such as abiotic stresses (salinity, drought, flooding, high temperatures etc.), plant-microbes and plant-pathogen interactions, and biostimulants (Fig. 1).

Figure 1. Untargeted metabolomics is the comprehensive analysis of all the measurable metabolites in a sample, including chemical unknowns, and targeted metabolomics, the measurement of defined groups of chemically characterized and biochemically annotated metabolites.
Noteworthy, recent advances in multivariate statistics, where supervised modelling approaches (e.g. partial least squares discriminant analysis or orthogonal projections to latent structures discriminant analysis, random forest algorithms) have integrated unsupervised statistics (like cluster analysis, k-means clusters, principal components analysis) allowing for proper data treatment thus providing more robust identification of differential metabolomic responses (Fig. 2). This, together with the availability of tools for data interpretation such as chemical enrichment, ontology or pathway analysis, has efficiently boosted the application of metabolomics in the field of agriculture including plant biostimulants (Fig. 3).
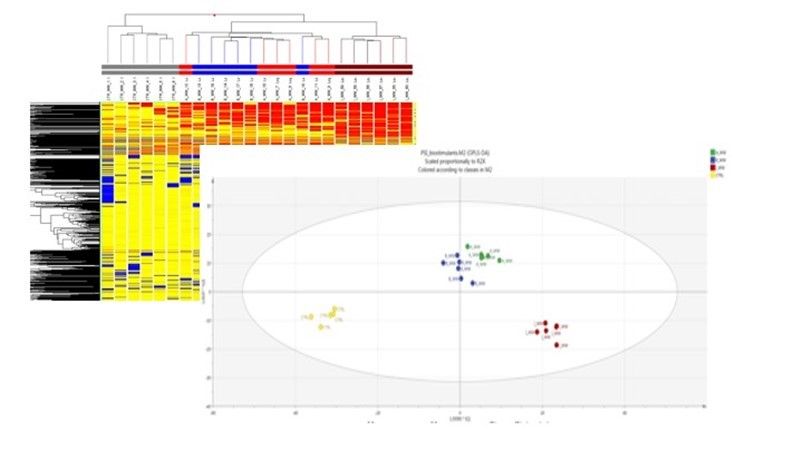
Figure 2. Unsupervised hierarchical cluster analysis carried out from metabolomic profiles following application of selected protein hydrolysates (upper figure). Score plot of Orthogonal Projection to Latent Structures Discriminant Analysis (OPLS-DA) supervised modeling carried out on metabolomic profiles following application of selected protein hydrolysates where the variation between groups was separated into predictive and orthogonal components (below figure).

Figure 3. Biosynthesis processes involved in tomato cuttings to foliar application of vegetal-derived protein hydrolysates and stem application of indole-butyric acid (IBA) using a basal quick-dip in a concentrated solution of IBA. The x-axis represents each set of subcategories while the y-axis corresponds to the cumulative fold-change in comparison with untreated control.
For example, metabolomics allowed identifying the increased tolerance to salinity imposed by a vegetal-derived protein hydrolysate in lettuce; the investigation pointed out that the mitigation of oxidative stress, accumulation of osmolytes and modulation of sterols and terpenes as specific mechanisms in the observed increased tolerance to salinity (Lucini et al., 2015).
Similarly, a biopolymer-based biostimulant of vegetal origin positively modulated root metabolic profile in melon, following its application. Brassinosteroids could be identified as pivotal compounds triggered by the biostimulant, causing a hormonal imbalance (involving abscisic acid, cytokinins, and gibberellin-related compounds) also in shoots (Lucini et al., 2018).
Looking at examples from microbial biostimulants, the differences in metabolomic fingerprints allowed identifying the improved tolerance to drought exerted by arbuscular mycorrhiza (AMF) in wheat (Bernardo et al., 2019). Notably, this study reported a clear AMF x cultivar interaction. Although the benefits of arbuscular colonization under water limiting conditions were more evident in bread wheat, sugars and lipids resulted to be positively modulated by AMF colonization. Interestingly, the modulation of metabolites related to oxidative stress and a fine tuning of phytohormones crosstalk were also evidenced, with the stimulation of brassinosteroids biosynthesis in root being particularly evident. In another work, we added further clues on the complex response of crops to root colonization, by investigating the different wheat root exudation patterns imposed by arbuscular mycorrhiza and Trichoderma atroviride (Lucini et al., 2019). This study evidenced that most of the differences could be ascribed to phenolic compounds and lipids, phytosiderophores and chelating acids, amino acids derivatives and hormones. Irrespective from the specific response, metabolomics pointed out that the interaction between beneficial fungi and wheat roots produced a complex response in terms of root exudation, the most being totally unexplored and possibly involving a further cascade of processes.
As it can be noted from these few examples, a wide diversity of mechanisms is underlying the mode of action of plant biostimulants, with the tuning of phytohormone crosstalk network being a common (in)direct pivotal process. To efficiently unravel these processes, it might be worthy to combine metabolomics with other assays, like physiological measurements (gas exchange, chlorophyll fluorescence etc.), enzyme activity and markers of oxidative stress like malondialdehyde (a lipid peroxidation marker). However, very promising outcomes are being produced when metabolomics is combined with phenomics. Indeed, the novel high-throughput plant phenotyping approaches can be used either to preliminary screen or to complement metabolomics output, thus corroborating the postulations produced from differential metabolites. As an example, a recent work on tomato treated with a vegetal-derived protein hydrolysate combined phenotyping and metabolomics confirmed drought mitigation effects, identifying that treated plants exhibited an improved tolerance to reactive oxygen species imbalance likely resulted from a coordinated action of radical scavengers, signaling compounds (salicylic acid and hydroxycinnamic amides), as well as reduced biosynthesis of tetrapyrrole coproporphyrins (Paul et al., 2019a). The same approach (high-throughput plant phenotyping combined with metabolomics) was successfully used to screen several vegetal-derived protein hydrolysates for biostimulant activity in tomato (Paul et al., 2019b)
Concluding, although metabolomics is the “newborn” among other omics, its potential to unravel the biochemical processes involved in plant response to different stimuli is clearly emerging. In the field of plant biostimulants, this option might represent a valuable tool to depict the complex mechanisms related to the definition of their mode(s) of action.
For more information, you are invited to request access to the following Research Articles:
Bernardo L., Carletti P., Badeck F.W., Rizza F., Morcia C., Ghizzoni R., Rouphael Y., Colla G., Terzi V., Lucini L. (2019). Metabolomic responses triggered by arbuscular mycorrhiza enhance tolerance to water stress in wheat cultivars. Plant Physiology and Biochemistry, 137, 203-212. https://doi.org/10.1016/j.plaphy.2019.02.007
Lucini L., Rouphael Y., Cardarelli M., Canaguier R., Kumar P., Colla G. (2015). The effect of a plant-derived biostimulant on metabolic profiling and crop performance of lettuce grown under saline conditions. Scientia Horticulturae, 182, 124-133. https://doi.org/10.1016/j.scienta.2014.11.022
Lucini L., Rouphael Y., Cardarelli M., Bonini P., Baffi C., Colla G. (2018). A vegetal biopolymer-based biostimulant promoted root growth in melon while triggering brassinosteroids and stress-related compounds. Frontiers in Plant Science, 9:472. https://doi.org/10.3389/fpls.2018.00472
Lucini L., Colla G., Miras Moreno M.B., Bernardo L., Cardarelli M., Terzi V., Bonini P., Rouphael Y. (2019). Inoculation of Rhizoglomus irregulare or Trichoderma atroviride differentially modulates metabolite profiling of wheat root exudates. Phytochemistry, 157, 158-167. https://doi: 10.1016/j.phytochem.2018.10.033
Paul K., Sorrentino M., Lucini L., Rouphael Y., Cardarelli M., Bonini P., Miras Moreno M.B., Reynaud H., Canaguier R., Trtílek M., Panzarová K., Colla G. (2019a). A combined phenotypic and metabolomic approach for elucidating the biostimulant action of a plant-derived protein hydrolysate on tomato grown under limited water availability. Frontiers in Plant Science 10:493. doi: 10.3389/fpls.2019.00493
Paul K., Sorrentino M., Lucini L., Rouphael Y., Cardarelli M., Bonini P., Reynaud H., Canaguier R., Trtílek M., Panzarová K., Colla G. (2019b). Understanding the biostimulant action of vegetal-derived protein hydrolysates by high-throughput plant phenotyping and metabolomics: a case study on tomato. Frontiers in Plant Science. 10:47. doi: 10.3389/fpls.2019.00047



